
BrainWarp is a tool that supports the automatic integration of large collections of data on a common reference space – known as a standard brain. It provides functionalities for collaborative quality assurance, and supports the subsequent integration of data in BrainBase.
BrainBase, BrainBaseWeb und BrainGazer support the anatomical, spatial or structural search for very large collections of confocal microscopy images of the brain of the fruit fly, fruit fly larva or zebrafish. Our patented data structures allow an intuitive search for corresponding expression patterns of neurons in tens of thousands of 3D images almost in real-time. BrainGazer and BrainBaseWeb also provide a variety of visual analytics and visualization tools for the data.
At present, there are four instances of software that support public data collections:
www.larvalbrain.org: Standard version of larval fruit fly brain (Drosophila melanogaster) with anatomical annotations and a collection of genetic lines. The data collection was developed and made available within the framework of a project promoted by the DFG (German Research Foundation) and the FWF (Austrian Science Fund) in collaboration with the University of Leipzig (Professor Andreas Thum), UCLA (Professor Hartenstein), RWTH Aachen (Professor Dorit Merhof) and VRVis (Dr Katja Bühler).
fruitfly.tefor.net and zebrafish.tefor.net: A collection of confocal microscopy images with neural structures provided by the TEFOR infrastructure.
https://implegacy.brainbase.at: Older version of BrainBaseWeb with a collection of over 15,000 3D images of VT lines in the adult fruit fly (Drosophila melanogaster), which have been generated by Barry Dickson at IMP in Vienna.
BrainTrawler – integrative analysis system for complex brain networks:
Together with neuroscientists at the Haubensak Lab at IMP Vienna and Boehringer-Ingelheim, we are developing BrainTrawler, a web-based framework for studying heterogeneous neurobiological data from mice and humans. Using spatial indexing, we are able to analyse gene expression data and complex brain networks in real time. Classifying the data to hierarchically organised anatomical structures makes it possible to search at different anatomical levels. Together with intuitive network visualization, iterative visual questions and quantitative information, BrainTrawler enables multimodal networks to be genetically dissected at local/global level within the spatial context. Further information can be found here.
License information and information about collaborating on a joint research project can be obtained on request. Please contact DI Dr. Gerd Hesina via products(at)vrvis.at!
More about the research work of our Biomedical Image Informatics research group.

On March 3, 2020, Katja Bühler, head of our Biomedical Image Informatics Group, was awarded with the renowned TU Women's Prize.

During the 40th anniversary celebration of the Austrian Computer Society (OCG) on June 9, 2015, Johannes Sorger was awarded this year's OCG sponsorship award.

The paper "Visual and Quantitative Analysis of Higher Order Arborization Overlaps for Neural Circuit Research" was awarded.

The web-based tool "neuroMAP" of VRVis and IMP received the Best Paper Award of the IEEE BioVis 2013.

Within the project Larvalbrain 2.0, a dynamic multi-scale multi-level atlas and data collection of structural, molecular, physiological, and behavioral results of Drosophila melanogaster larvae will be established.
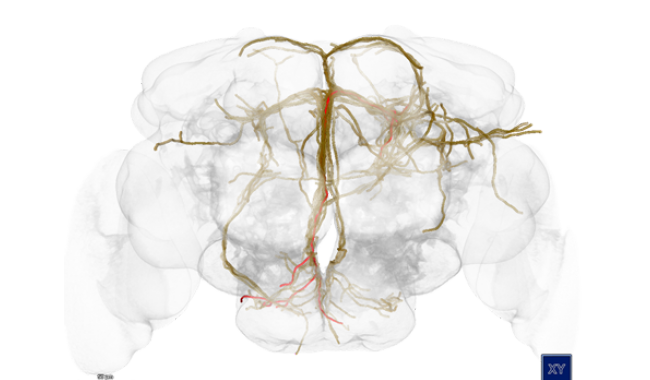
Understanding how the brain works is one of the biggest challenges addressed by neuroscientists today. Modern neuroscience research is extremely data-intensive and requires special software infrastructures to enable and accelerate the discovery of the complex interplay of genes, structure and function.

VRVis contributes data analytics and visualization tools tailored to support and accelerate research of the Haubensak Group at the Institute of Molecular Pathology Vienna.

For centuries, neuroscientists have been mapping the brain. Until now, the step from simple maps to a generally accepted model has proved to be extremely difficult. In this project, a 4D atlas of the brain of the fruit fly larva is being built.
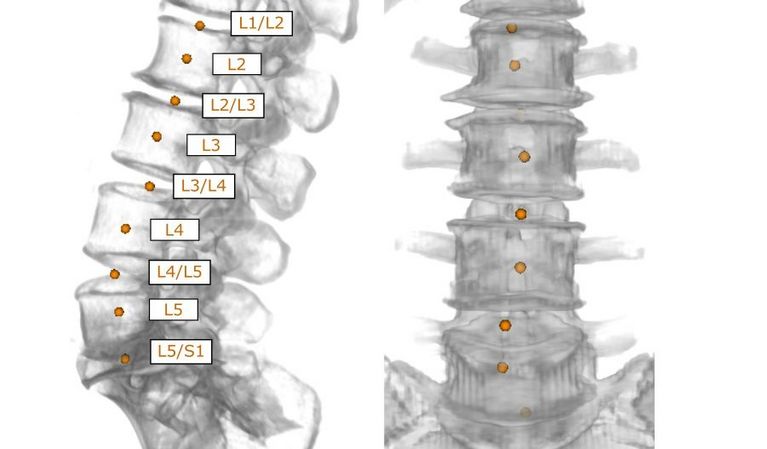
The long-term vision of this applied research project is to use available data resources to improve image-based diagnostics based on complex data in daily clinical routine.
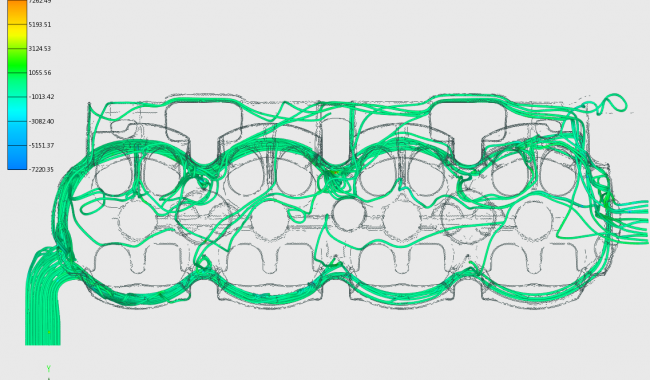
The strategic project forms the organizational and scientific hub for the realization of an area wide integrative visual computing approach. It covers joint strategic research and development on fundamental challenges in all application projects.